We currently have genome sequences for over two dozen yeast species, which occupy a wide range of ecological niches. By comparing these genomes we can learn about the basic evolutionary processes at the level of genes. (This paper is a nice but technical introduction to the subject; the Genolevures website is one of the best online sources of information about yeast evolution.)
I came across an interesting paper about budding yeast that discusses a specific instance of adaptive evolution in stunning detail - the alteration of galactose utilization pathway in several yeast species that have adapted to different environments. As the authors of the paper state,
"We have found that at least three independent lineages of yeast have inactivated or lost most or all of the genes of the GAL pathway while leaving interacting genes intact. The parallel losses of this entire pathway are extreme examples of an emerging general principle that gene-inactivation events reveal specific functions affected by recent changes in the ecological pressures acting on a species."
Let's back up and explain what the galactose utilization pathway is: Yeast metabolize the sugar galactose (see the picture below) through several steps mediated by specific enyzmes that convert galactose into a modified form of glucose, which is then further metabolized to generate high energy molecules and byproducts like alcohol or lactic acid. This is a well characterized pathway, meaning that the specific proteins involved and their functions are known, as are the modifcations made to the galactose molecule. In rough outline, here is the galactose utilization pathway: The protein Gal2p allows galactose into the cell, where it is transformed in sequence by the proteins Gal1p, Gal7p, and Gal5p. By this point the galactose molecule has been changed around to become a modified form of glucose, which can then be further metabolized. This pathway is tightly regulated by a set of control proteins (such as Gal4p), which act to turn the synthesis of the pathway proteins on or off.
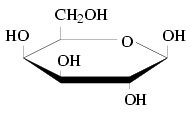
A yeast species that lives in an environment where galactose is a major food source needs a functioning galacose utilization pathway. Species that adapt to a difference niche, where galactose utilization is not essential, would be expected to lose the genes involved in this pathway, because those genes would no longer be preserved by natural selection.
Now, back to the paper:
The authors looked for the GAL genes in the genomes of four yeast species that cannot metabolize galactose, and found that in three of these species, all of the dedicated GAL genes were completely missing, with on exception:
The yeast species E. gossypii harbors a form of the GAL4 gene, but the DNA sequence of this gene suggests that it has undergone rapid evolutionary change, possibly an indication that this GAL4 gene has been recruited for a new function in E. gossypii. This is an excellent hypothesis begging to be tested - one could delete this gene in E. gossypi and look at the consequences.
[Note that I'm skipping over a lot of technical detail - determining that this particular gene has undergone rapid DNA sequence change involves some statistical testing - you can find more details here. The point is, when we say that a gene has undergone rapid evolutionary change, it involves more than just looking at a genes sequence and saying "yep, looks like evolution to me."]
These three yeast species which were missing any trace of most of the GAL genes were species that broke off early from the evolutionary branch leading to baker's yeast. They lost their ability to use galactose a long time ago, and the relentless erosion by mutation in the absence of selection has erased the GAL genes from their genomes. (See the figure below - the species that can't use galactose are listed in red, and the red vertical bars represent points in time where each lineage lost the ability to use galactose)
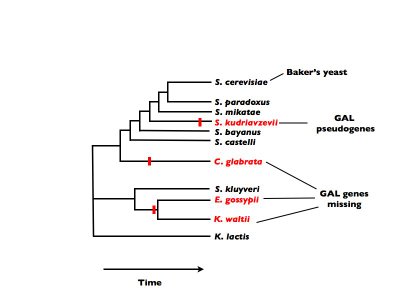
The species S. kudriavzevii however, is much more closely related to baker's yeast. It cannot use galactose, even though all of its close species cousins still can. This indicates that it lost the ability to use galactose more recently. As a result, the non-functional remnants of the GAL genes have not yet been erased from its genome - they are all still present as pseudogenes. (Pseudoegenes are genes that look like protein-coding genes, except they have been rendered non-functional by some inactivating mutation.)
S. kudriavzevii appears to have recently adapted to a new ecological niche - unlike other close relatives of baker's yeast, S. kudriavzevii is not found in sugar-rich environments; it was isolated from soil and decaying leaves. As a result of this change in lifestyle, there was no longer any selective pressure to maintain the GAL genes, thus they are slowly being erased by mutation.
To sum up:
1) 4 yeast species among the set of 11 studied cannot use galactose for metabolic energy. Each of these 4 species has lost its functional GAL genes, while species that can use galactose have retained their GAL genes.
2) Species that lost the ability to use galactose a long time ago have had nearly all traces of their GAL genes erased by the relentless force of mutation over time (we're on a scale of 100 million years or more). In one species, the GAL4 gene appears to have been recruited for a new (as yet unknown) function.
3) One species lost its ability to use galactose more recently (the last few million years); in this species non-functional GAL genes are still present as pseudogenes.
4) This is a fascinating example of gene inactivation resulting from the relaxation of the force of natural selection that occurs as organisms adapt to a new environment.
The real bottom line: What we find when we look at genomes is absolutely unexplainable unless we accept the bedrock tenets of evolution: 1) today's diverse life forms diverged from common ancestors, and 2) mutation and natural selection are major driving forces of evolutionary change.




1 comment:
Post a Comment